Running inference tools
As machine learning (ML) becomes more popular in HEP analysis, coffea also provide tools to assist with using ML tools within the coffea framework. For training and validation, you would likely need custom data mangling tools to convert HEP data formats (NanoAOD, PFNano) to a format that best interfaces with the ML tool of choice, as for training and validation, you typical want to
have fine control over what computation is done. For more advanced use cases of data mangling and data saving, refer to the awkward array manual and uproot/parquet write operations for saving intermediate states. The helper tools provided in coffea focuses on ML
inference, where ML tool outputs are used as another variable to be used in the event/object selection chain.
Why these wrapper tools are needed
The typical operation of using ML inference tools in the awkward/coffea analysis tools involves the conversion and padding of awkward array to ML tool containers (usually something that is numpy-compatible), run the inference, then convert-and-truncate back into the awkward array syntax required for the analysis chain to continue. With awkward arrays’ laziness now being handled entirely by `dask <https://dask-awkward.readthedocs.io/en/stable/gs-limitations.html>`__, the conversion
operation of awkward array to other array types needs to be wrapped in a way that is understandable to dask. The packages in the ml_tools package attempts to wrap the common tools used by the HEP community with a common interface to reduce the verbosity of the code on the analysis side.
Example using ParticleNet-like jet variable calculation using PyTorch
The example given in this notebook be using `pytorch <https://pytorch.org/>`__ to calculate a jet-level discriminant using its constituent particles. An example for how to construct such a pytorch network can be found in the docs file, but for mltools in coffea, we only support the TorchScript format files to load models to ensure operability when scaling to clusters. Let us first start by downloading the example ParticleNet model file and a small
PFNano compatible file, and a simple function to open the PFNano with and without dask.
[1]:
!wget --quiet -O model.pt https://github.com/CoffeaTeam/coffea/raw/master/tests/samples/triton_models_test/pn_test/1/model.pt
!wget --quiet -O pfnano.root https://github.com/CoffeaTeam/coffea/raw/master/tests/samples/pfnano.root
[2]:
from coffea.nanoevents import NanoEventsFactory
from coffea.nanoevents.schemas import PFNanoAODSchema
def open_events():
factory = NanoEventsFactory.from_root(
{"file:./pfnano.root": "Events"},
schemaclass=PFNanoAODSchema,
)
return factory.events()
Now we prepare a class to handle inference request by extending the mltools.torch_wrapper class. As the base class cannot know anything about the data mangling required for the users particular model, we will need to overload at least the method prepare_awkward:
The input can be an arbitrary number of awkward arrays or dask awkward array (but never a mix of dask/non-dask array). In this example, we will be passing in the event array.
The output should be single tuple
a+ single dictionaryb, this is to ensure that arbitrarily complicated outputs can be passed to the underlyingpytorchmodel instance likemodel(*a, **b). The contents ofaandbshould benumpy-compatible awkward-like arrays: if the inputs are non-dask awkward arrays, the return should also be non-dask awkward arrays that can be trivially converted tonumpyarrays via aak.to_numpycall; if the inputs are dask awkward arrays, the return should be still be dask awkward arrays that can be trivially converted via ato_awkward().to_numpy()call. To minimize changes to the code, a simpledask_awkward/awkwardswitcherget_awkward_libis provided, as there should be (near)-perfect feature parity between the dask and non-dask arrays.In this ParticleNet-like example, the model expects the following inputs:
A
Njets x2coordinate x100constituents “points” array, representing the constituent coordinates.A
Njets x5feature x100constituents “features” array, representing the constituent features of interest to be used for inference.A
Njets x1mask x100constituent “mask” array, representing whether a constituent should be masked from the inference request.
In this case, we will need to flatten the
Eevents xNjets structure, then, we will need to stack the constituent attributes of interest viaak.concatenateinto a single array.
After defining this minimum class, we can attempt to run inference using the __call__ method defined in the base class.
[3]:
from coffea.ml_tools.torch_wrapper import torch_wrapper
import awkward as ak
import dask_awkward
import numpy as np
class ParticleNetExample1(torch_wrapper):
def prepare_awkward(self, events):
jets = ak.flatten(events.Jet)
def pad(arr):
return ak.fill_none(
ak.pad_none(arr, 100, axis=1, clip=True),
0.0,
)
# Human readable version of what the inputs are
# Each array is a N jets x 100 constituent array
imap = {
"points": {
"deta": pad(jets.eta - jets.constituents.pf.eta),
"dphi": pad(jets.delta_phi(jets.constituents.pf)),
},
"features": {
"dr": pad(jets.delta_r(jets.constituents.pf)),
"lpt": pad(np.log(jets.constituents.pf.pt)),
"lptf": pad(np.log(jets.constituents.pf.pt / jets.pt)),
"f1": pad(np.log(np.abs(jets.constituents.pf.d0) + 1)),
"f2": pad(np.log(np.abs(jets.constituents.pf.dz) + 1)),
},
"mask": {
"mask": pad(ak.ones_like(jets.constituents.pf.pt)),
},
}
# Compacting the array elements into the desired dimension using
# ak.concatenate
retmap = {
k: ak.concatenate([x[:, np.newaxis, :] for x in imap[k].values()], axis=1)
for k in imap.keys()
}
# Returning everything using a dictionary. Also perform type conversion!
return (), {
"points": ak.values_astype(retmap["points"], "float32"),
"features": ak.values_astype(retmap["features"], "float32"),
"mask": ak.values_astype(retmap["mask"], "float16"),
}
# Setting up the model container
pn_example1 = ParticleNetExample1("model.pt")
# Running on dask_awkward array
dask_events = open_events()
dask_results = pn_example1(dask_events)
print("Dask awkward results:", dask_results.compute()) # Runs file!
/Users/saransh/Code/HEP/coffea/.env/lib/python3.11/site-packages/coffea/ml_tools/helper.py:175: UserWarning: No format checks were performed on input!
warnings.warn("No format checks were performed on input!")
Dask awkward results: [[0.0693, -0.0448], [0.0678, -0.0451], ..., [0.0616, ...], [0.0587, -0.0172]]
For each jet in the input to the torch model, the model returns a 2-tuple probability value. Without additional specification, the torch_wrapper class performs a trival conversion of ak.from_numpy of the torch model’s output. We can specify that we want to fold this back into nested structure by overloading the postprocess_awkward method of the class.
For the ParticleNet example we are going perform additional computation for the conversion back to awkward array formats:
Calculate the
softmaxmethod for the return of each jet (commonly used as the singular ML inference “scores”)Fold the computed
softmaxarray back into nested structure that is compatible with the original events.Jet array.
Notice that the inputs of the postprocess_awkward method is different from the prepare_awkward method, only by that the first argument is the return array of the model inference after the trivial from_numpy conversion. Notice that the return_array is a dask array.
[4]:
class ParticleNetExample2(ParticleNetExample1):
def postprocess_awkward(self, return_array, events):
softmax = np.exp(return_array)[:, 0] / ak.sum(np.exp(return_array), axis=-1)
njets = ak.count(events.Jet.pt, axis=-1)
return ak.unflatten(softmax, njets)
pn_example2 = ParticleNetExample2("model.pt")
# Running on dask awkward
dask_events = open_events()
dask_jets = dask_events.Jet
dask_jets["MLresults"] = pn_example2(dask_events)
dask_events["Jet"] = dask_jets
print(dask_events.Jet.MLresults.compute())
/Users/saransh/Code/HEP/coffea/.env/lib/python3.11/site-packages/dask_awkward/lib/structure.py:901: UserWarning: Please ensure that dask.awkward<count, npartitions=1>
is partitionwise-compatible with dask.awkward<divide, npartitions=1>
(e.g. counts comes from a dak.num(array, axis=1)),
otherwise this unflatten operation will fail when computed!
warnings.warn(
[[0.528, 0.528, 0.524, 0.523, 0.521, 0.52, 0.519, 0.519], ..., [0.528, ...]]
Of course, the implementation of the classes above can be written in a single class. Here is a copy-and-paste implementation of the class with all the functionality described in the cells above:
[5]:
class ParticleNetExample(torch_wrapper):
def prepare_awkward(self, events):
jets = ak.flatten(events.Jet)
def pad(arr):
return ak.fill_none(
ak.pad_none(arr, 100, axis=1, clip=True),
0.0,
)
# Human readable version of what the inputs are
# Each array is a N jets x 100 constituent array
imap = {
"points": {
"deta": pad(jets.eta - jets.constituents.pf.eta),
"dphi": pad(jets.delta_phi(jets.constituents.pf)),
},
"features": {
"dr": pad(jets.delta_r(jets.constituents.pf)),
"lpt": pad(np.log(jets.constituents.pf.pt)),
"lptf": pad(np.log(jets.constituents.pf.pt / jets.pt)),
"f1": pad(np.log(np.abs(jets.constituents.pf.d0) + 1)),
"f2": pad(np.log(np.abs(jets.constituents.pf.dz) + 1)),
},
"mask": {
"mask": pad(ak.ones_like(jets.constituents.pf.pt)),
},
}
# Compacting the array elements into the desired dimension using
# ak.concatenate
retmap = {
k: ak.concatenate([x[:, np.newaxis, :] for x in imap[k].values()], axis=1)
for k in imap.keys()
}
# Returning everything using a dictionary. Also take care of type
# conversion here.
return (), {
"points": ak.values_astype(retmap["points"], "float32"),
"features": ak.values_astype(retmap["features"], "float32"),
"mask": ak.values_astype(retmap["mask"], "float16"),
}
def postprocess_awkward(self, return_array, events):
softmax = np.exp(return_array)[:, 0] / ak.sum(np.exp(return_array), axis=-1)
njets = ak.count(events.Jet.pt, axis=-1)
return ak.unflatten(softmax, njets)
pn_example = ParticleNetExample("model.pt")
# Running on dask awkward arrays
dask_events = open_events()
dask_jets = dask_events.Jet
dask_jets["MLresults"] = pn_example(dask_events)
dask_events["Jet"] = dask_jets
print(dask_events.Jet.MLresults.compute())
print(dask_awkward.necessary_columns(dask_events.Jet.MLresults))
[[0.528, 0.528, 0.524, 0.523, 0.521, 0.52, 0.519, 0.519], ..., [0.528, ...]]
{'from-uproot-3196a0c383555cda3738c112acd1c70e': frozenset({'nJetPFCands', 'PFCands_dz', 'nPFCands', 'Jet_eta', 'Jet_nConstituents', 'PFCands_phi', 'PFCands_d0', 'nJet', 'PFCands_pt', 'JetPFCands_pFCandsIdx', 'PFCands_eta', 'Jet_phi', 'Jet_pt'})}
In particular, analyzers should check that the last line contains only the branches required for ML inference; if there are many non-required branches, this may lead the significant performance penalties.
As per other dask tools, the users can extract how dask is analyzing the processing the computation routines using the following snippet.
[6]:
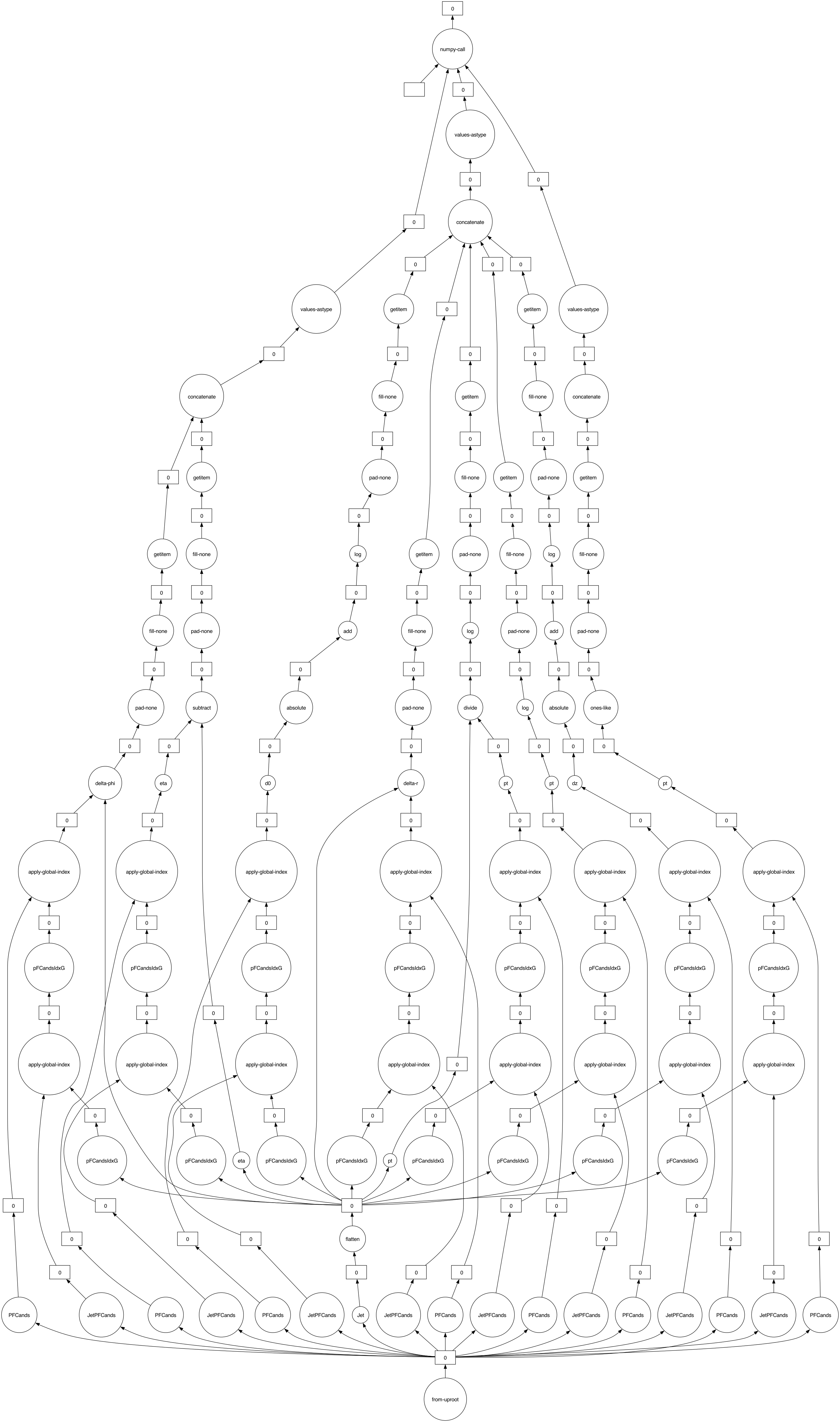
print(dask_results.dask)
dask_results.visualize(optimize_graph=False)
HighLevelGraph with 104 layers.
<dask.highlevelgraph.HighLevelGraph object at 0x29169cc10>
0. from-uproot-3196a0c383555cda3738c112acd1c70e
1. JetPFCands-dd2ea51f30214bf71538143d483f24f9
2. PFCands-bc578074fd7542d617f1a321b55033b8
3. JetPFCands-2022a279fa9f32fb5958ee0196c7bc9c
4. PFCands-83b1509b3ea29e972a2c83951cb53cb6
5. JetPFCands-c3abed82cbd768736fc7d2efe53b1bfb
6. PFCands-1082ee2cc592b1a0c1b8219ddbb9df76
7. JetPFCands-95b391cea3695b0e90f6ff4136821900
8. PFCands-448b56417f7e6e78f111bda34eb8ba7e
9. JetPFCands-31a2eb013adf67227780245e9f6e7654
10. PFCands-80d89bc6034885fc4a99e252f4c76d87
11. JetPFCands-fa15abc1502f6fa51ba0d6608dac9af8
12. PFCands-4b547af5e660141b1c5163448bc75e50
13. JetPFCands-65dd6ed1fed0463a350740a761960f79
14. PFCands-18a47bdc7f8227a81fc30fa63c20e0b8
15. JetPFCands-a30dac67296389d5ee6ed32d038d9a29
16. PFCands-16b38f51395d73298b304b5b74560b87
17. Jet-2a79d0b5a69da035a6f63a34642205aa
18. flatten-645563137107a3dabf8c0252326c099b
19. pFCandsIdxG-be948845416432cfc7843dc1818979ba
20. apply-global-index-5d46f157ea0ed464f14667860b7f9fa0
21. pFCandsIdxG-3658836fb2e3ea21bc533848c17dbf9c
22. apply-global-index-7ea60f265dccb1ecc1371332dda18513
23. pt-e2b62bfe096c321d604a32fbf89668b2
24. ones-like-d424640d3caa1d220630da9879f9a6d4
25. pad-none-61123924df10b8c261bbc98bc5c2b24f
26. fill-none-16100aa7cabf6e0f37054b3cda2d9d7e
27. getitem-fad74ed67d4b95383a9d0afde0e454b4
28. concatenate-axisgt0-0615838b6257bb1dd9e0ad365df899a7
29. values-astype-2f87bb6f27689b84fbd8379d88089848
30. pFCandsIdxG-91d61065f2be395629a7f7c2f4c75a4c
31. apply-global-index-0920d60e59835d92a5dacb96a448cb0d
32. pFCandsIdxG-a081c6286992ba2fccaa2f2a11518923
33. apply-global-index-e509f72064969e949a238db289c45072
34. dz-1730bacc99456e5ba3253a49df42e172
35. absolute-e01baff37424469f75445b90028366da
36. add-a70004667fa4b10107c6efb1f2a97989
37. log-a3603e25871acedaabcc3e2099d4bcb2
38. pad-none-4a4dedef54637f16ef2b271c1475f31a
39. fill-none-c5a035e8e95e0c9e361ff39d1ebd9f2d
40. getitem-bc1fb20d7b9e5fbc6a51534ccd3054f3
41. pFCandsIdxG-d6802f98055e2806250a9c3227728372
42. apply-global-index-e35fa5ba3d7597da70c007766683a812
43. pFCandsIdxG-632a8052988ed5a2a8938a273ab2d333
44. apply-global-index-b5e8f87704f5d545cf817d2762944d25
45. d0-29e79e5b5a7de17a03a3d475ff89c599
46. absolute-c8d09ca24f3086d1e1e2b9414b4ed022
47. add-8761d4cf88a177af713d19002a764f4f
48. log-ddae64d2513ac33f2660b46d6854a3dc
49. pad-none-d886677ded46c3f4e8d87456e6e680e2
50. fill-none-d45149e4990e4da381d842d166e7c5ba
51. getitem-836d4f0d5523ecbd8d6db1e90ae2b3b6
52. pt-10e04c1e9f951b6eea81cb85c498833a
53. pFCandsIdxG-f039caebcf4c11cb6e22e91d73d58061
54. apply-global-index-b7778b7df69d732f785a0b8c9d57ca7c
55. pFCandsIdxG-32fb1e5b834c5af2cfb38c19c71f7901
56. apply-global-index-75fa94598ada38bc2c7ac256829aba69
57. pt-35cd28a2662d400d82c8b0e0bf1043be
58. divide-b70f48381fa00780673100850a77be64
59. log-59467169551d668c177b279d7ce41e08
60. pad-none-d5344cecf02aace568fb6048ed540975
61. fill-none-82e8626d15b47f726ea132d0ce2172db
62. getitem-4404eac5ca31322afe93d2f586df1bc7
63. pFCandsIdxG-f0b011a50db292fb2b871686ee0a4ca4
64. apply-global-index-adabd8aa56e51de7d7da71bbc78e54d7
65. pFCandsIdxG-61b55e563c2ef8da75531d37f4588e46
66. apply-global-index-4ddf80eb0996d4e224663e3718eea052
67. pt-54e90537bdb3fe376c28a70cd79127a6
68. log-8784340a02993c5f7a0a94affb9303b7
69. pad-none-8e9507bad08a1e98574503d852bb8e08
70. fill-none-35f0eb9a5380d8e9b99d3fdd92720c63
71. getitem-3122b07919ce90a875c60fe3379baaf2
72. pFCandsIdxG-6751177f307689956c9a5195ee32bf1c
73. apply-global-index-7d0b6c07be47f20e367642fb8e283891
74. pFCandsIdxG-5057f535f83a536bfa670cd1be195413
75. apply-global-index-35c83043c8ca14ca9330f4e462ee80d3
76. delta-r-a4e12fdba83dc2391ced6c43b1f899fd
77. pad-none-529c1a1cfc95ddd8615681183fb06572
78. fill-none-ea30a23fe8ebad07ad2326697bc04680
79. getitem-9f44943e437f2bb256ccc100cc97f2da
80. concatenate-axisgt0-b5c7c2098dc5ed82427a7f31eb5ed39a
81. values-astype-e8e2df120704dbe38f61b0a4b0263819
82. pFCandsIdxG-aa02ecbc510ba5db2b1bec3d1007c8a7
83. apply-global-index-149b1ad33ead558e3b736d22c3a261fc
84. pFCandsIdxG-988a4723504c8a86b25fce7b6dcd1ed0
85. apply-global-index-ac5938c59513f8973b8a0cc39f69be2a
86. delta-phi-f7ff1ff2df14e7932e2b711fb13b15ab
87. pad-none-4fbed45948badcca50ce362c653eefff
88. fill-none-1f7d59ea6f5c76a6b81dbf2e358271e1
89. getitem-0dc813c8eeffc021339d7e91776fa416
90. pFCandsIdxG-e798cc4121bd681e903080f4f1389924
91. apply-global-index-d7b8b6b56eeeb86c9683ebc761346f24
92. pFCandsIdxG-f6807458d1b798c1765dc82431de0630
93. apply-global-index-97d92fe2282fbbc36a1d400cebc3f8d6
94. eta-f24b1fc33dca394d0f6803cc7784e37e
95. eta-d97bf469e0213a29c85425c1e3d91b04
96. subtract-6138b5e5f850f64d920c23efa39303ca
97. pad-none-9b25781acc91d8b6517bb37dae719dd0
98. fill-none-d0965fb8ec4099be0707b78aecaf4a1b
99. getitem-9f41b44078a5993adc37fbadf25ee227
100. concatenate-axisgt0-fae11878826f65e15ee7d9eb1e0043d7
101. values-astype-bf0acc24ca1d686bc7e4dc91eee546e8
102. ParticleNetExample1-d4d79650-ea96-4f0d-9187-9a87f15fe12c
103. numpy-call-ParticleNetExample1-906b63a30d0298bea4410f9b6ff1d666
[6]:
Or a peek at the optimized results:
[7]:
dask_results.visualize(optimize_graph=True)
/Users/saransh/Code/HEP/coffea/.env/lib/python3.11/site-packages/coffea/ml_tools/helper.py:175: UserWarning: No format checks were performed on input!
warnings.warn("No format checks were performed on input!")
[7]:
Comments about generalizing to other ML tools
All ML wrappers provided in the coffea.mltools module (triton_wrapper for triton server inference, torch_wrapper for pytorch, and xgboost_wrapper for xgboost inference) follow the same design: analyzers is responsible for providing the model of interest, along with providing an inherited class that overloads of the following methods to data type conversion:
prepare_awkward: converting awkward arrays tonumpy-compatible awkward arrays, the output arrays should be in the format of a tupleaand a dictionaryb, which can be expanded out to the input of the ML tool likemodel(*a, **b). Notice some additional trivial conversion, such as the conversion to available kernels forpytorch, converting to a matrix format forxgboost, and slice of array fortritonis handled automatically by the respective wrappers. To handle both dask/non-dask arrays, the user should use the providedget_awkward_liblibrary switcher.postprocess_awkward(optional): converting the trivial converted numpy array results back to the analysis specific format. If this is not provided, then a simpleak.from_numpyconversion results is returned.
If the ML tool of choice for your analysis has not been implemented by the coffea.mltools modules, consider constructing your own with the provided numpy_call_wrapper base class in coffea.mltools. Aside from the functions listed above, you will also need to provide the numpy_call method to perform any additional data format conversions, and call the ML tool of choice. If you think your implementation is general, also consider submitting a PR to the coffea repository!